AskOmics is a collaborative web platform for data integration and query using the semantic web (RDF and SPARQL).
This guide is intended for users who want to analyze their project-specific data with AskOmics. Throughout the guide, you will find
To do this tutorial, you will need an AskOmics instance. you can use genouest instance. You will also need input data. Data are available here.
Account creation and management¶
Login or signup into AskOmics¶
AskOmics is a mutli-user plateform. To use it, you will need an account on the instance. Use the
Hands-on
Create your AskOmics account (or login with your existing one)
Once your are logged, you can use all the functionalities of AskOmics.
Manage your account¶
To manage your account, use the
Uses the forms to change your personal information.
Data integration¶
AskOmics convert project specific data into RDF triples automatically. It can convert CSV/TSV, GFF and BED files. It can also integrate RDF data.
Data upload¶
The first step is to upload the input files into AskOmics. Go on the Files page by clicking on
You can upload files from your computer, or distant files using an URL.
Hands-on
Upload the files TAIR1_GFF3_genes_mRNA.gff, transcriptomics.tsv and qtl.tsv from your computer into AskOmics
Uploaded files are displayed into the files table.
Next step is to convert this files into RDF triples. This step is called Integration. Integration will produce a RDF description of your data: the Abstraction.
Integration¶
The integration convert input files into RDF triples, and load them into an RDF triplestore. AskOmics can convert CSV/TSV, GFF3 and BED files. Select input files from the Files page, and click on the
CSV/TSV¶
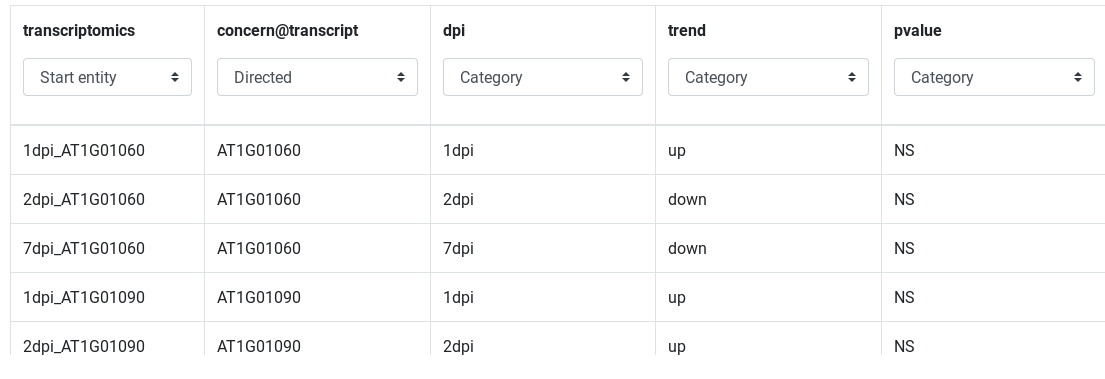
CSV/TSV files are previewed on the integration page as a table. The first column describe the entity to integrate, other columns describe relations or attributes of this entity. Each column type is automatically detected by AskOmics, but it can be overrided.
- The first columns can be an Entity or an Start entity (default). Only a Start entity can be a start point of a query.
- Columns that respect the format
relation@entitydescribe a relation of the current entity to another entity.relationis the name of the relation andentityis the targeted entity. Relations can be Directed (from the current entity to the target one) or Symetric (both direction). - Other columns can be Attributes. They can be Numeric, Text or Category.
Warning
The Category type store all the different values of the category in the abstraction. Use it only for attributes that have a limited number of different values
- Other columns can also be Faldo attributes. Faldo attributes describe genetic location of an entity. a FALDO entity will be converted in RDF using the FALDO ontology.
Hands-on
Integrate transcriptomics.tsv and qtl.tsv. a QTL is a locatable entity. Set chromosome, start and end as FALDO attributes (reference, start and end). The transcriptomics entity contain a relation to a gene entity (not integrated yet). Other attributes have repeated values. They can be integrated as category
GFF¶
GFF files contain genetic coordinate of entities. Each entities contained in the GFF file are displayed on the preview page. Select the entities you want to integrate.
Hands-on
Integrate the TAIR1_GFF3_genes_mRNA.gff file. The GFF contain the locatable entities gene and mRNA. Select both and integrate them. The gene entities is the entity targeted by transcriptomics.
BED¶
BED contain also locatable entities. You have to specify an entity name since the BED format don't specify it.
RDF¶
RDF file can be directly integrated into AskOmics. This RDF file have to contain the abstraction and the data to be correctly used in AskOmics.
Manage integrated datasets¶
All integrated files are stored in a specific named graph in the RDF triplestore. The named graph obtained are called datasets. You can manage the integrated datasets on the
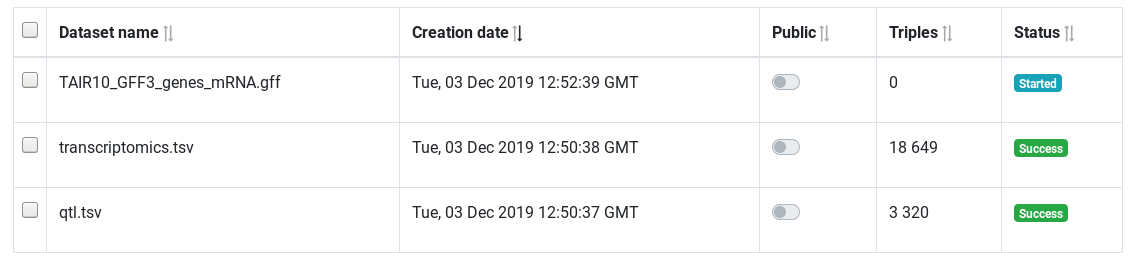
The table show all integrated datasets. The status column show if the datasets is fully integrated or in the process of being integrated. You can delete datasets independently.
Query¶
Once your data are integrated, the time has come to make requests on these data. Go to the Ask page by clicking on
Start point¶
On the Ask page, all entities available are showed on the page.
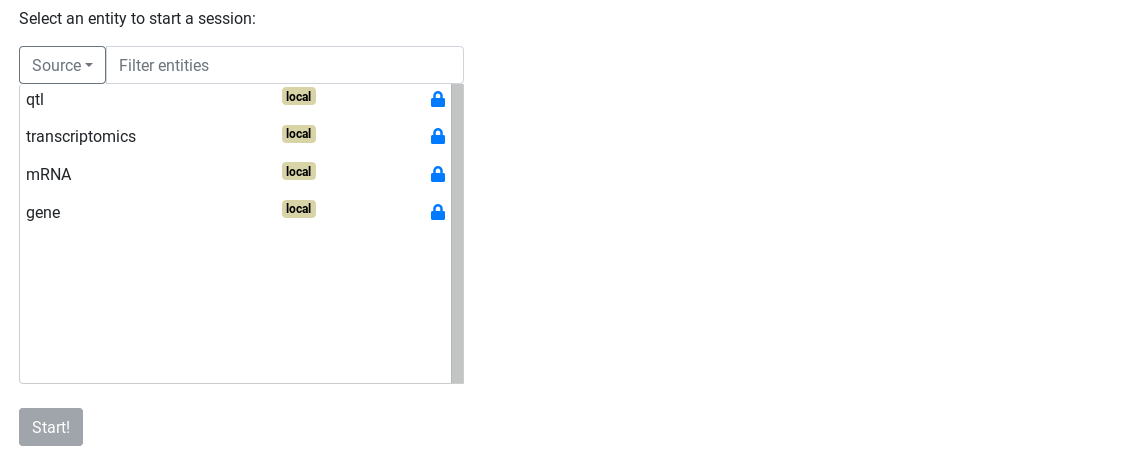
The badge
If the entities are numerous, you can filter them by enter a string in the Filter entities field, or by select the source of the entities. Here, we only have local entities so we can only filter with text.
To start a query, select a start point. You must choose the start point according to what you want to obtain at the end of the query. For example, you want genes that are on chromosome 3, the start point to choose is gene.
Start points can be filtered with the Filter entities box. a badge show where the entity is. Here, we only have local entities.
Simple query¶
To start a query, select a start point and click on
Hands-on
Start a query with gene
The query builder is divided into two side. The left side is the entity graph and the right side is the attribute view.
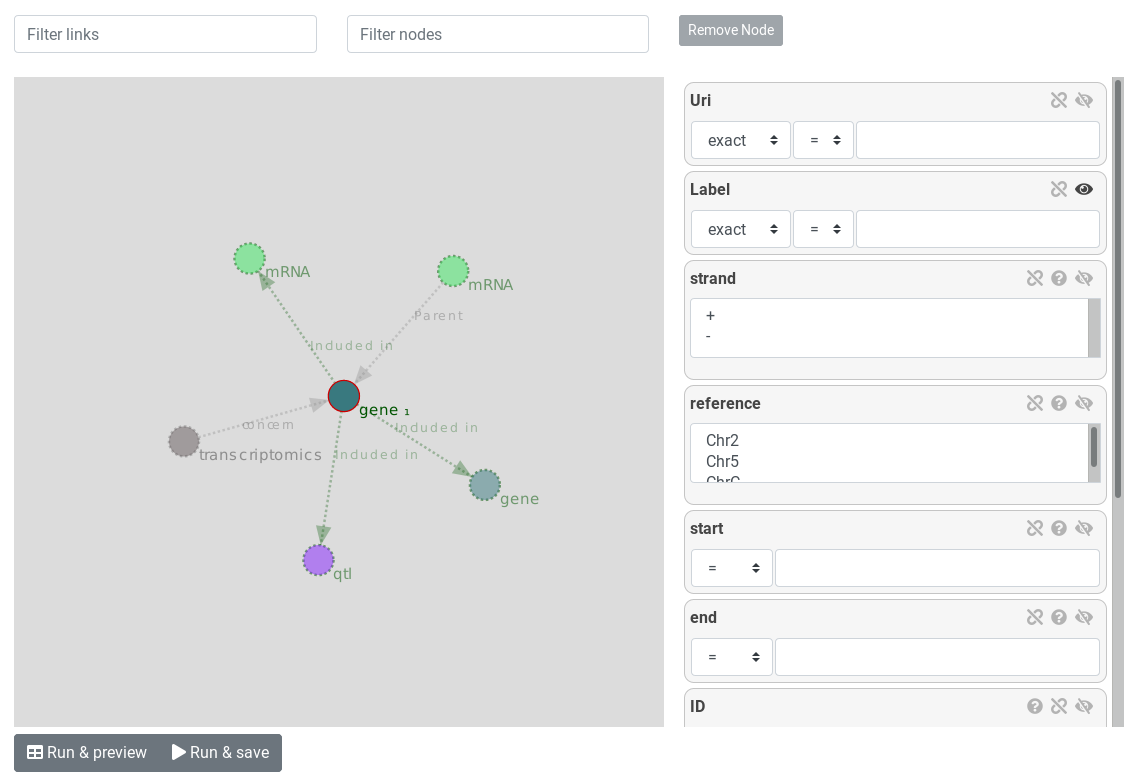
On the entity graph, entities are represented as bubbles and relations are represented as arrows. We can see the gene start point selected (surrounded by a red circle). From the selected gene bubble start relations to other entities. This relations are suggested (transparent and dotted). On the attribute view, attributes of the selected entities are displayed. At this point, the represented query is "Give me all genes".
Hands-on
Launch a preview of the query by clicking on
A preview of the results are displayed on the bottom of the page.
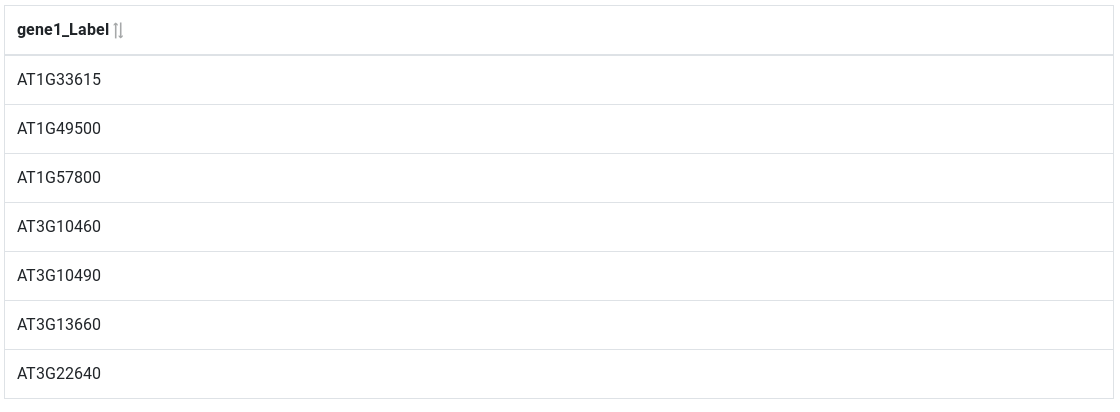
Complex query¶
To build a complex query, we need to apply constraints to our node. Two kinf of constraints can be apply. Constraints on relations and constraints on attributes. This constraints are called filters From this point, we will construct the following query:
All genes that are significantly over-expressed on day post-infection one (dpi1) and under-expressed at dpi 3 and 7.
Filters on attributes¶
On the attribute view, you see all attributes of the gene entity. Each attribute is represented as an attribute box. Depending of the type of attribute (numeric, text or category), Each box type have common properties and specific properties.
- Show/hide
Each attribute have a show/hide button, represented with an eye icon . By default, only the attribute label is showed (), all other attributes are hidden () Click on the icon to display an attribute.
Hands-on
Show the reference, start and end of the gene entity, and preview the results
The common properties are the show/hide button, the
- Text attribute

A text attribute can be filtered by a string or a regexp. Use filter types to change the filter from exact match to regexp and to change the match to a not match.
Warning
exact match is the most efficient filter. regexp and negative match can take much more time.
Hands-on
Filter the gene label with AT1G06820 and preview the result. Then, perform a regexp filter with ^AT1G06
- Numeric attribute
Numeric attribute can be filtered with a number. The filter type ise used to compare the number with =, ≠, >, <, ≤ or ≥.
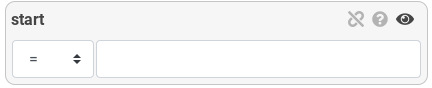
Hands-on
Filter the start attribute with values superior or equal to 2000000 and preview results
- Category attribute
A category is an attribute who have a limited number of values. Each values are showed on the attribute box. To filter a category, click on each value. use ctrl + click to select several values or remove them.
Hands-on
Select the reference Chr1 and Chr2 and preview results
At this point, we have "all genes whose label start with AT1G06 located on chr1 and chr2, and with a start value superior or equal to 2000000".
Filter on relations¶
Filter on relations allow to link an entity to other entities. On the entity graph, the selected entity is surrounded by a red circle, and other entities are proposed (transparent and dotted). To instantiate a entity linked to the selected one, click on it.
Hands-on
Return to the gene. Then, instantiate a transcriptomics entity, linked to our gene entity by clicking on it and preview results
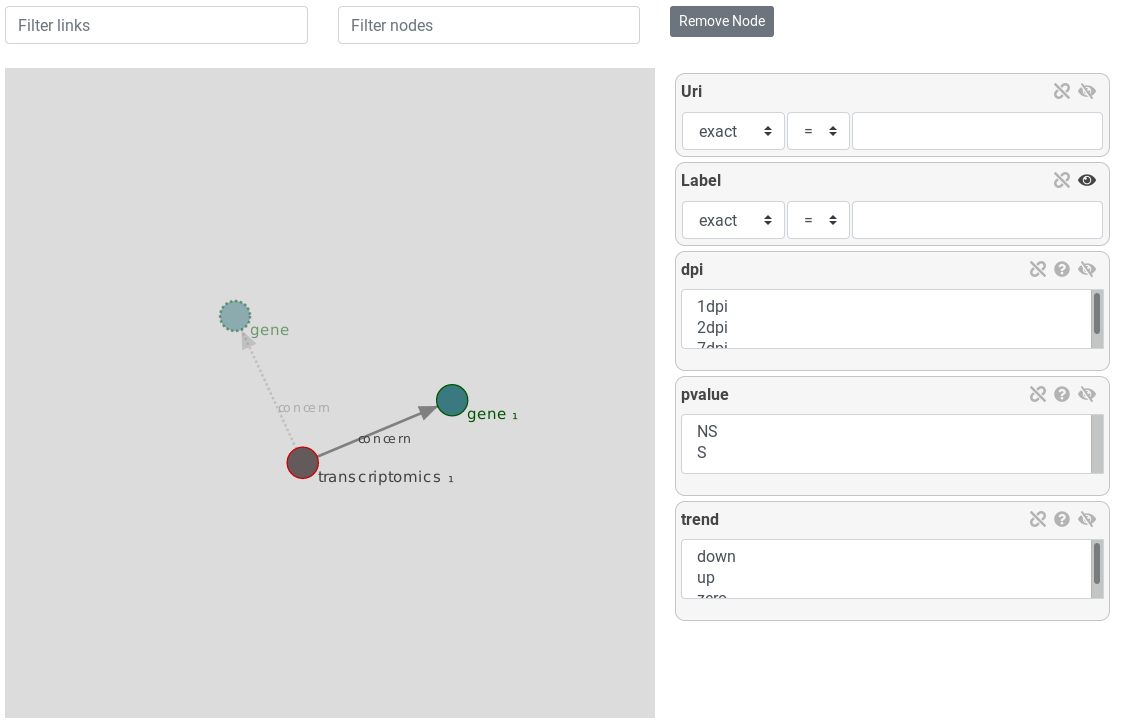
This query represent all genes linked to a transcriptomics experiment. The transcriptomics entity represent transcriptimics experiments ...
On the attributes view, values can be filtered to contraints the results.
Hands-on
Filter the transcriptomics entity to get all experiments at day post infection one (dpi) with synificative values (S) and with an up trend
At this step, we have genes that are synificatively over-expressed at day one post infection.
Several same instance of entities can be linked. To get other condition, we can instanciate other transcriptomics experiments to gene.
Hands-on
Return to gene, and instanciate a new transcriptomics entity. Filter it with dpi2, and a down trend. Then re-return to gene and do the same to a third transcriptomics entity, but a dpi7. Preview results
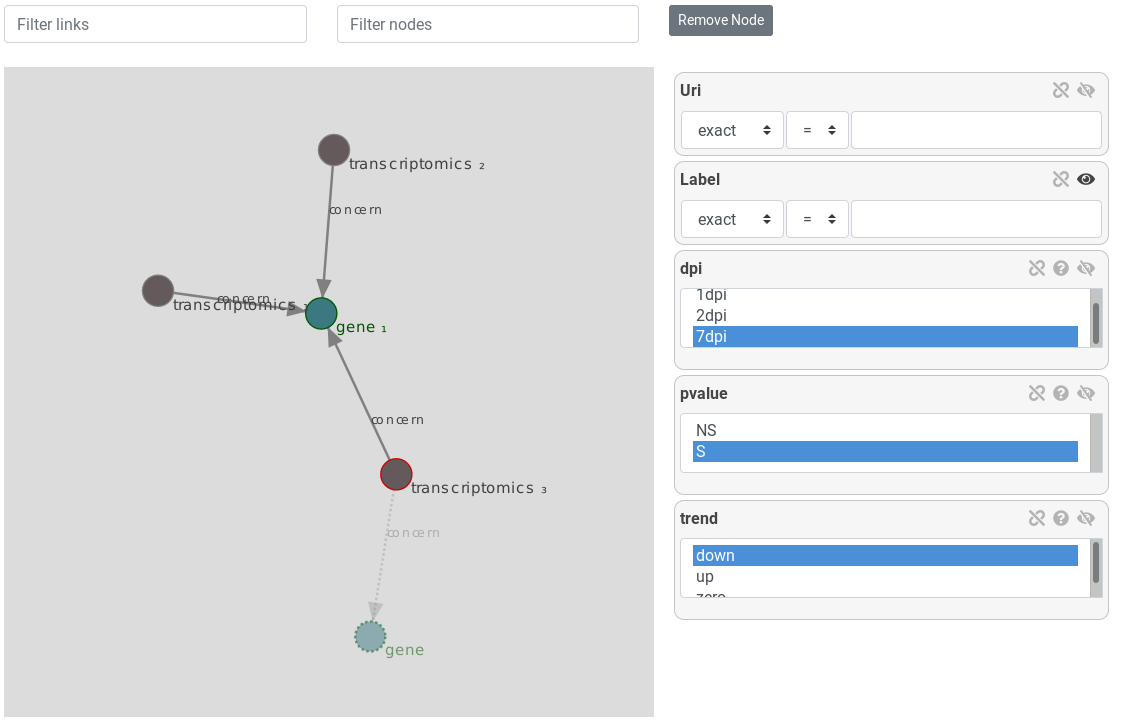
Now, results show genes that are synificatively over-expressed at day one and under-expressed at day 2 and 7.