Galaxy is a scientific workflow, data integration, and data and analysis persistence and publishing platform that aims to make computational biology accessible to research scientists that do not have computer programming or systems administration experience.
AskOmics can be used with a Galaxy instance in two way:
- With a dedicated AskOmics, import Galaxy datasets into AskOmics and export AskOmics results into Galaxy.
- In Galaxy: use AskOmics Interactive Tool inside Galaxy
Link AskOmics with Galaxy¶
Create a Galaxy API key¶
On your Galaxy account, go to the top menu User → API Keys and copy your API key. Yhis API key is unique identifier that will be used for AskOmics to access to data.
Enter Galaxy API key into your AskOmics account¶
On AskOmics, got to

Once a Galaxy account is added to AskOmics, you can access to all your Galaxy Datasets from AskOmics.
Upload a file from Galaxy¶
On the
Send result and query to Galaxy¶
On the
- Send result to Galaxy: Send the result file to the last recently used history
- Send query to Galaxy: send the json graph state that represent the AskOmics query
Import a saved query from Galaxy¶
On the
Galaxy AskOmics Interactive Tool¶
Galaxy Interactive Tools (GxITs) are a method to run containerized tools that are interactive in nature into the Galaxy interface. AskOmics have his GxIT available into several instances:
Launch AskOmics IT¶
Search for the AskOmics Interactive tool using the search bar.
Choose input files to automatically upload them into AskOmics
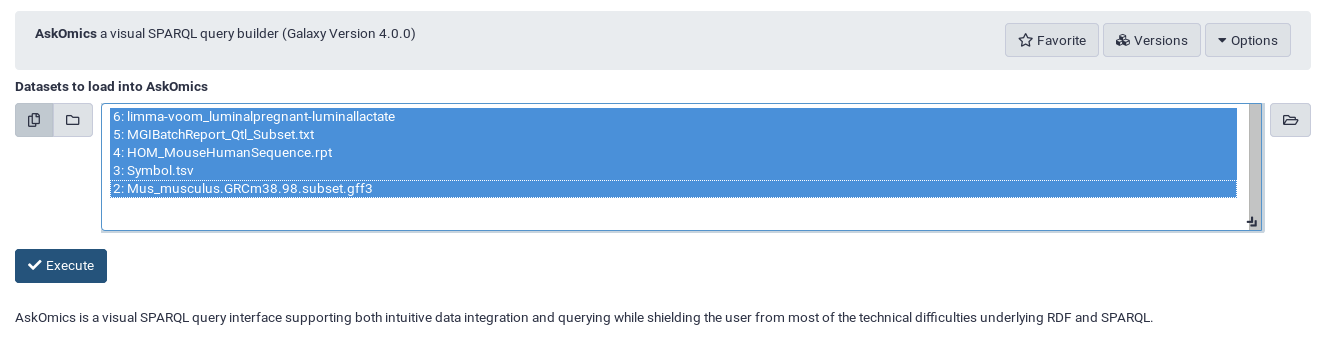
Tip
You will able to add more input files later
A dedicated AskOmics instance will be deployed into the Cluster. Wait few minutes and go to the instance using the click here to display link.
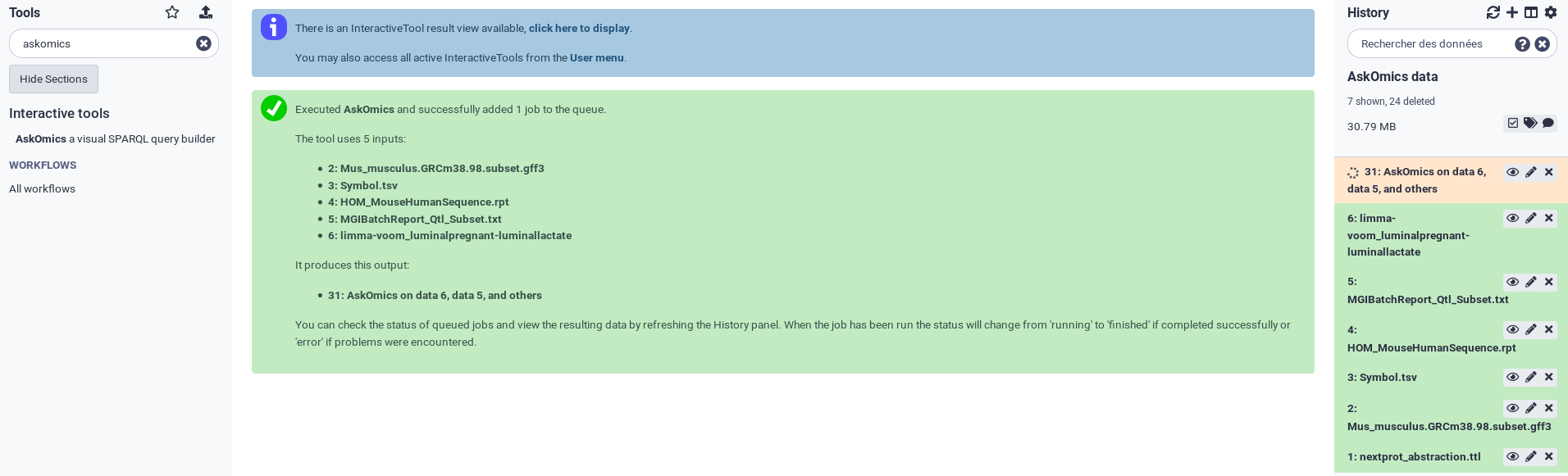
Once you are into your AskOmics instance, you can see your uploaded files into the
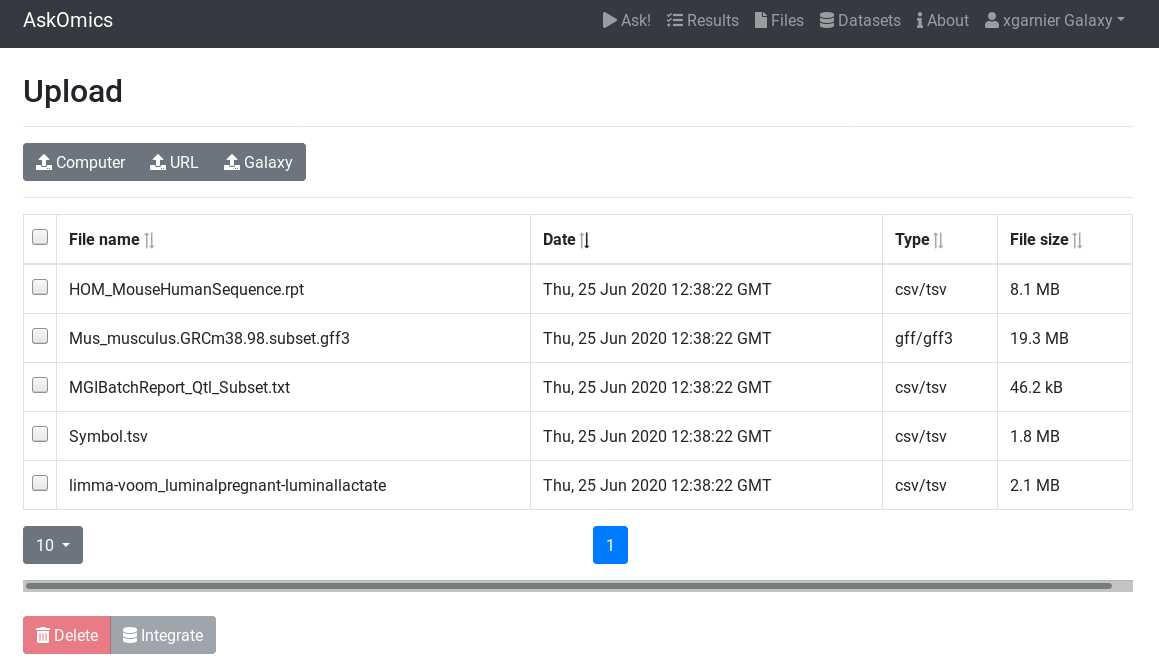
Upload additional files¶
in addition to the
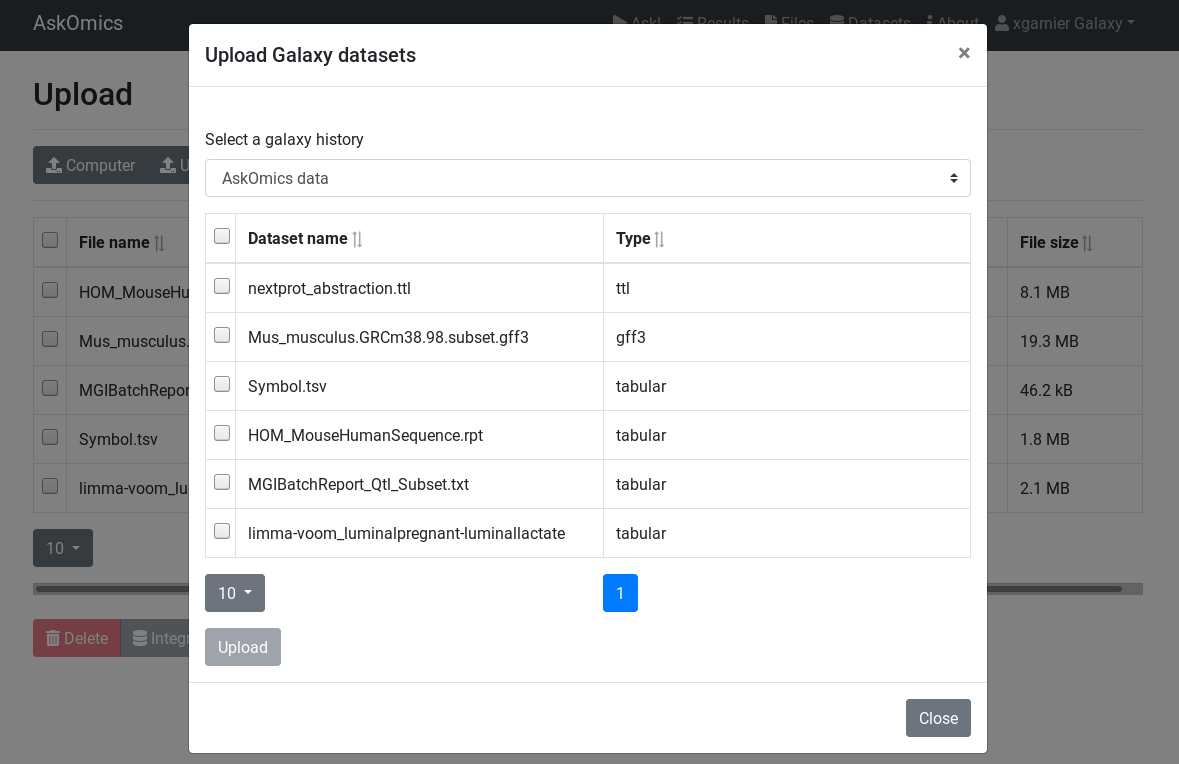
Integrate and Query¶
follow the tutorial to integrate and query your data.
Export Results into your Galaxy history¶
Once you have your result, Use the Send result to Galaxy to export a TSV file into your last recently used Galaxy history.