A federated query is a query who involve several SPARQL endpoints. AskOmics have his dedicated endpoint for the integrated data, but it is also possible to query external resources.
Define an external endpoint¶
The first step is to define an external endpoint. External endpoint have their own description. To Display external entities, AskOmics need the Abstraction of the distant endpoint. This external abstraction can be build automatically or manually.
Auto-generate external abstraction with abstractor
Abstractor is a command line tool that auto-generate an abstraction from a distant endpoint.
pip install abstractor
abstractor -e <endpoint_url> -p <entity_prefix> -o <output_file>
Warning
Abstractor scan all things in the SPARQL endpoint. You may review the generated file to delete unwanted things.
Integrate external abstraction into AskOmics
Once external endpoint's abstraction is generated, its time to add it into AskOmis. Upload it and integrate it. During integration, use advanced options and specify the external endpoint.
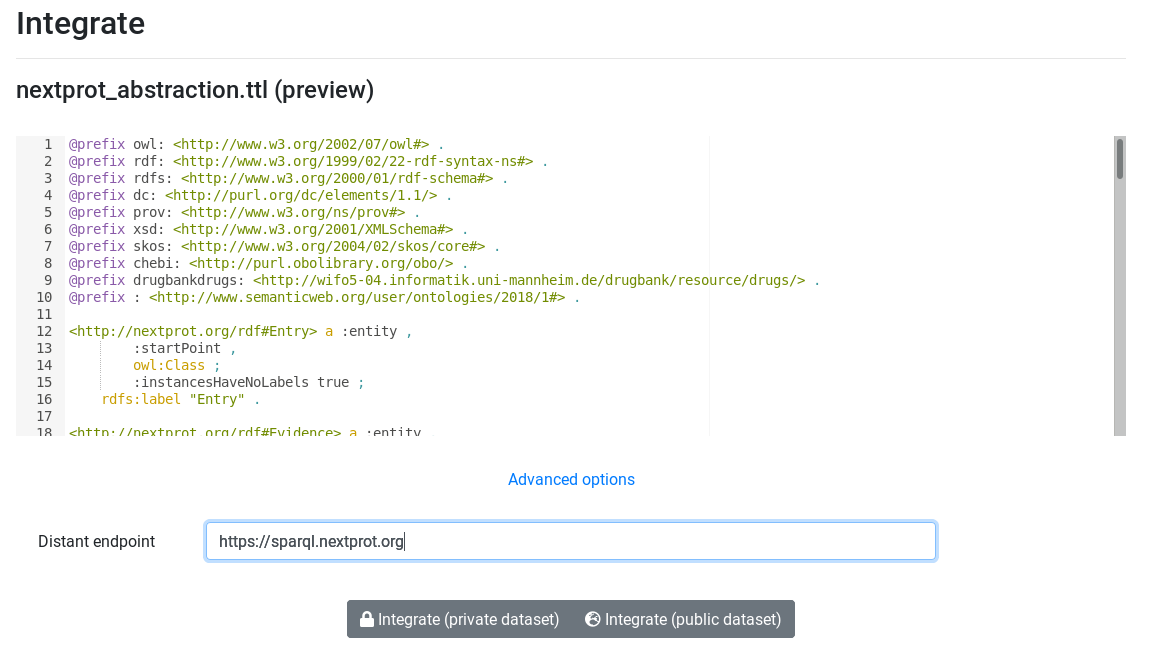
Query external endpoint¶
Simple query
External startpoint are not displayed by default on the start page. Use the Source dropdown button to display external entities.
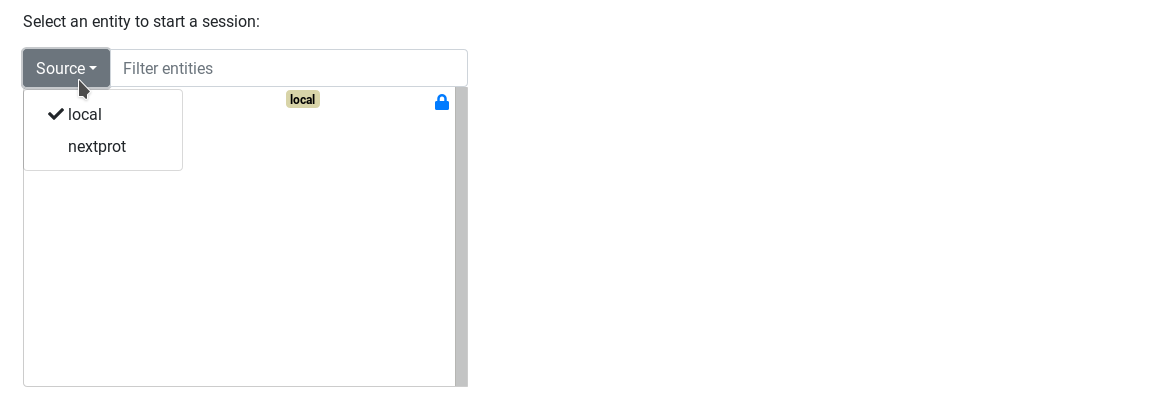
Federated query
External entities can be interrogate just as local entities. But to link a local dataset to the external endpoint, the file must be structured in a certain way.
Build file
The input file must describe the relation with the external entity. It goes through the header, who must contain the URI of the targeted entity. Content of the file must also be the exact uri of the targeted entity.
For example, the file below describe en entity gene who is linked to an external entity Gene. The external one is prefixed with the full uri used in the external endpoint. In the content of the file, full URI have to be used to.
| gene | value | concern@http://nextprot.org/rdf#Gene |
|---|---|---|
| gene_1 | 0 | http://nextprot.org/rdf/gene/ENSG00000169594 |
| gene_2 | 1 | http://nextprot.org/rdf/gene/ENSG00000156603 |
Perform a federated query
Once the relations are well described, link between local and distant entities are automatically done by AskOmics. The Query is distributed to the concerned endpoint and results are returned like a classic query.